What is NeST?
The NeST (Nested Systems in Tumors) map is a hierarchy of 395 protein systems that are recurrently mutated in one or more cancer types. It provides a resource of cancer mechanisms under selection for somatic mutations and captures canonical cancer pathways and protein complexes, along with novel protein assemblies on which mutations unexpectedly converge.
NeST was assembled by creating an Integrated Protein Network combining evidence of interactions of five major data types: physical protein-protein interactions (novel and previously known), mRNA co-expression, protein co-expression, sequence similarity, and genetic co-dependence. Multiscale community detection was then applied to the network to detect protein communities (a.k.a. "systems") at different size resolutions. Communities detected at small size resolutions can overlap with one another and nest within communities at larger size resolutions, resulting in a hierarchy of systems. Finally, a statistical model called HiSig was developed to identify the systems on which cancer mutations significantly converge, yielding the NeST Systems Map of 395 systems.
Accessing NeST
The NeST Systems Map is accessible via the NDEx (Network Data Exchange), an online commons for biological networks. In NDEx, the map appears as a hierarchical network of nodes (protein systems) and edges (hierarchical containment of one system by another). Node color corresponds to the number of tumor types in which a system is recurrently mutated, and node size is proportional to the system size (number of proteins). The map can also be directly downloaded to the Cytoscape network visualization and analysis platform. The NeST model is also available in OBO format on Zenodo.
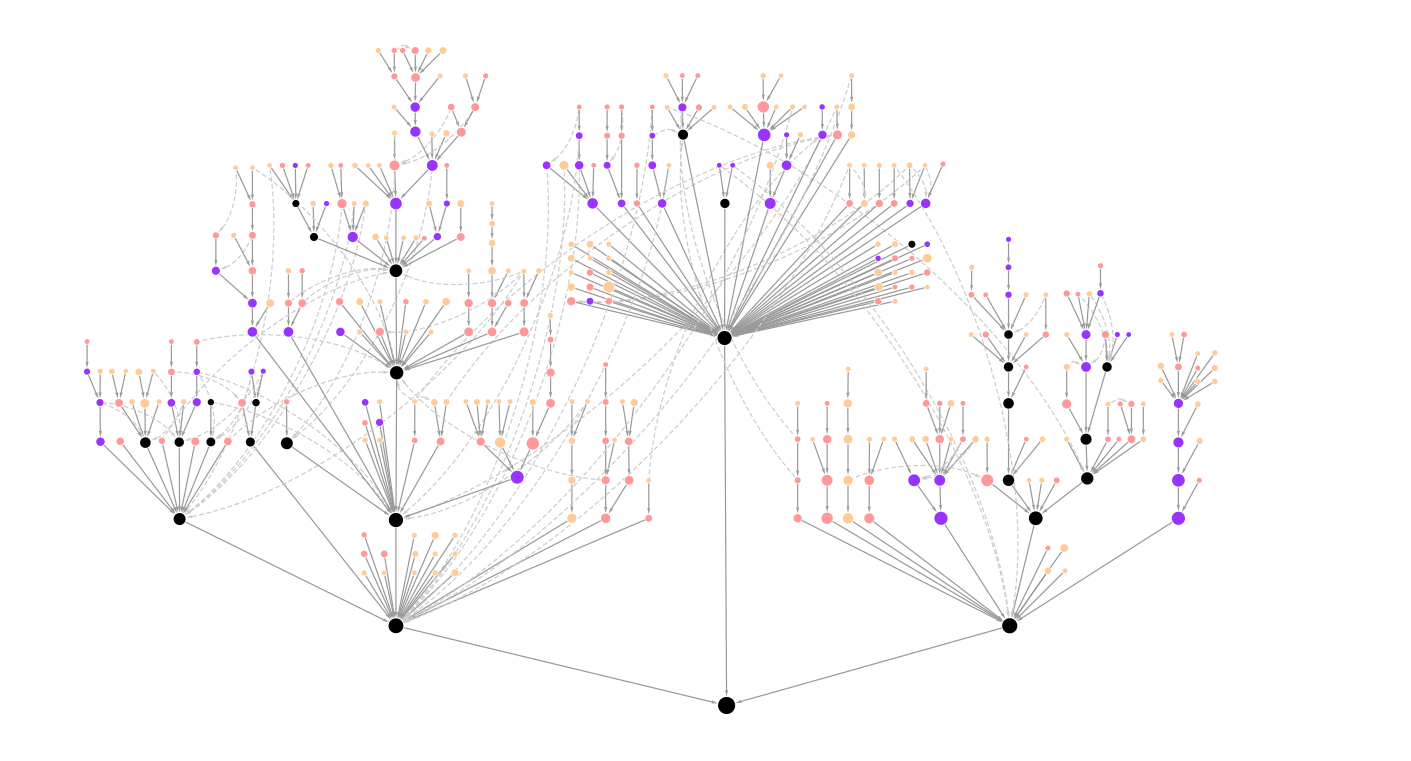
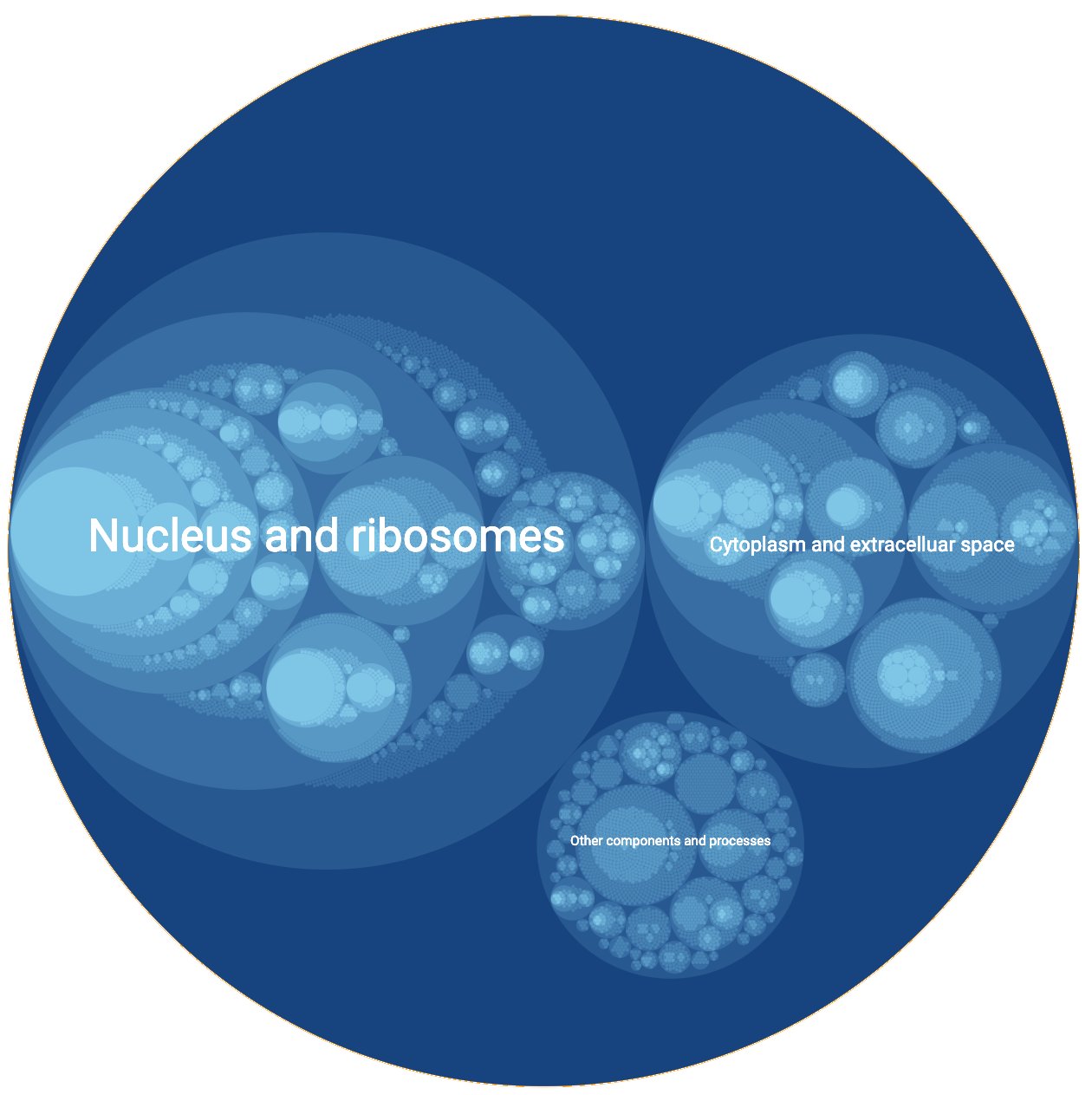
An alternative means of exploring the NeST map is provided by HiView, in which the hierarchical structure is represented as a kaleidoscopic collection of nested circles. Each circle represents a protein system. If one system contains another, the latter appears as a nested circle inside the former. HiView allows the user to (1) interactively zoom across many scales in the model, from an expansive view of the entire map to more focused views of particular subsystems; (2) browse the network data supporting the inference of each system; and (3) search for genes and systems based on their names.
Availability of Code
The bioinformatics tools used to create the NeST map are open-source and can be reused or adapted as a template for hierarchical modeling of other diseases influenced by complex genetic alterations:
CliXO: multiscale community detection
HiSig: identifying significantly mutated systems
How to cite
Zheng F*, Kelly MR*, Ramms DJ, Heintschel ML, Tao K, Tutuncuoglu B, Lee JJ, Ono K, Foussard H, Chen M, Herrington KA, Silva E, Liu SN, Chen J, Churas C, Wilson N, Kratz A, Pillich RT, Patel DN, Park J, Kuenzi B, Yu MK, Licon K, Pratt D, Kreisberg JF, Kim M, Swaney DL, Nan X, Fraley SI, Gutkind JS, Krogan NJ†, Ideker T†.Interpretation of cancer mutations using a multiscale map of protein systems. Science. 2021 Oct;374(6563):eabf3067. doi: 10.1126/science.abf3067. Epub 2021 Oct 1. PMID: 34591613 *Equal Contribution, †Co-Corresponding Authors
